TLDR: Scaling up pandas is hard. With Modin, we took a first-principles approach to parallelizing the pandas API. Rather than focus on implementing what we knew was easy, we developed a theoretical basis for dataframes—the abstraction underlying pandas—and derived a dataframe algebra that can express the 600+ pandas operators in under 20 algebraic operators.
The pandas dataframe library is widely used by millions of users to perform a variety of data-centric tasks spanning transformation, validation, cleaning, exploration, and featurization. The popularity of the library stems from the flexible tabular data model underlying dataframes, as well as the rich set of supported functions. Compared to relational databases, pandas is more flexible, expressive, and convenient.
However, this flexibility, expressiveness, and convenience comes at a cost. pandas is often non-interactive on moderate-to-large datasets, and breaks down completely when operating on datasets beyond main memory. This is due to multiple reasons. First, pandas is single threaded, meaning that it cannot leverage multiple cores in a machine or cluster. Second, pandas operates entirely in memory and is inefficient in so – leading to disruptive out-of-memory errors.
To address these shortcomings, we developed Modin, a parallel dataframe system as a drop-in replacement for pandas. Modin has had over 2.5M downloads, 75 contributors across 12+ institutions, and more than 6.8k GitHub stars (as of Feb 2022). In designing and developing Modin, we set out to address pandas scalability challenges without changing any pandas behavior and functionality – a challenge some might have previously thought impossible.
Try Ponder Today
Start running pandas and NumPy in your database within minutes!
So why is parallelizing pandas so hard?
At a fundamental level, optimizing 600+ functions directly is exceedingly difficult: the sheer engineering effort would be back-breaking. Moreover, these functions can be chained in arbitrary ways, leading to an exponential number of possibilities that need to be optimized individually.


In contrast, consider relational databases. Query processing and optimization for databases have been around for more than 40 years. The reason why database queries in SQL are easy to parallelize is because there are only a small handful (less than 10) of relational operators: filters, projections, group bys, and joins, among others. It took the database community over four decades to perfect the optimization and parallelization of each relational operator, but now, we can demonstrate that relational databases are truly scalable.
If it took the database community 40 years to perfect 10 operators, imagine how long it would take to perfect all of the 600+ functions in pandas. On top of this, there are a lot of differences between the relational and dataframe data models that make this even more difficult! Since we didn’t have 40 years (or more!) to wait, we needed a new approach.

Our key insight: deriving and parallelizing the “core” operators
As we studied the pandas API and pandas usage, we concluded that the massive pandas API can be represented by less than 20 core operators. These operators can be flexibly applied along either axis: rows, columns, and even individual cells. For example, `min` accepts an input axis argument that specifies whether to return the minimum value along rows or columns. (Note that this is unlike relational operators that can only be applied along one axis. For example, filters are only along rows.)
The dataframe algebra operators include low-level operators such as map, explode, and reduce, relational operators such as group-by, join, rename, sort, concat, and filter, metadata manipulation operators such as infer_types, filter_by_types, to/from_labels, linear algebra operators such as transpose, and order-based operators such as window and mask.
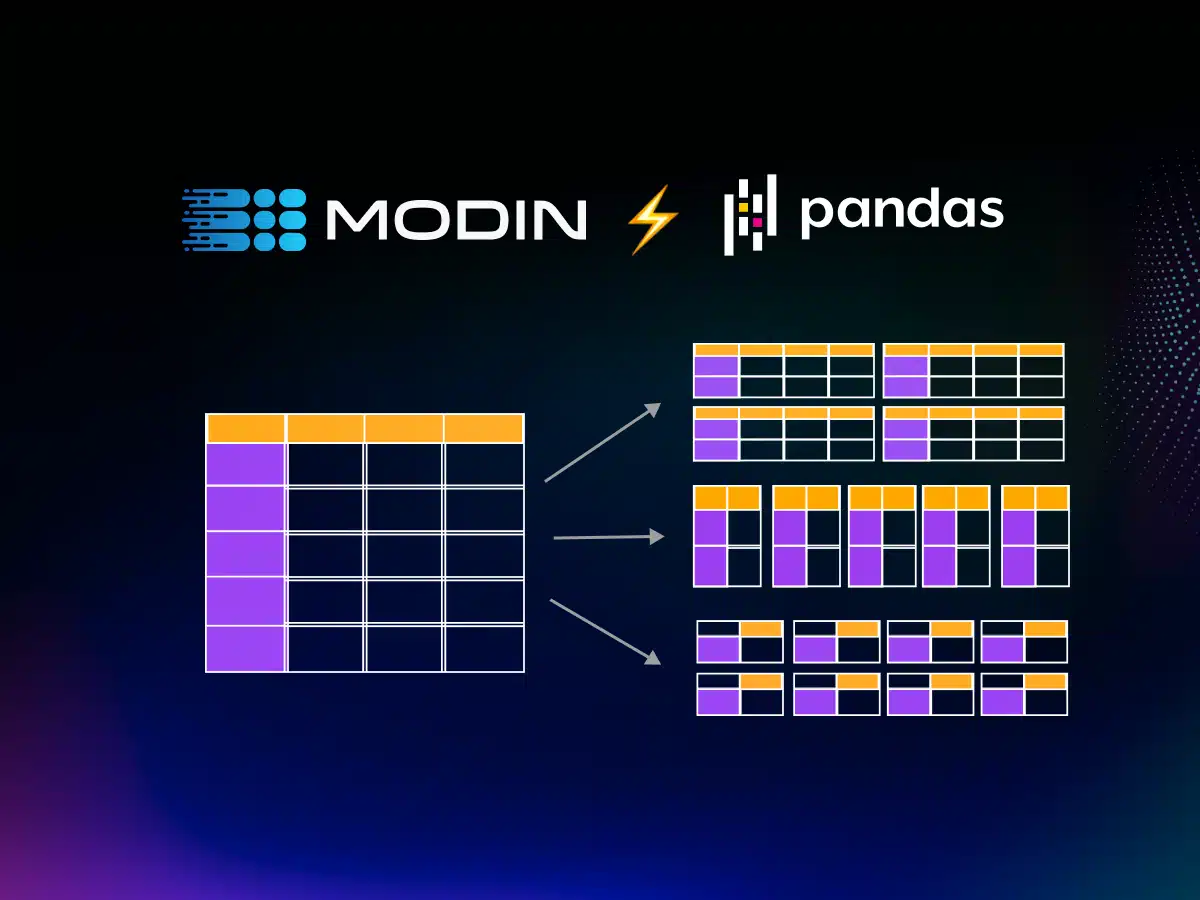
Now that we had identified the what of dataframes (i.e. what they can do), we were ready to explore the how (i.e. how to parallelize). We developed a set of decomposition rules that allow us to rewrite the core operators into a series of smaller operations that can be applied in parallel across data partitions, vertical, horizontal, or block-based, depending on the operation. For example, a `min` operation that is applied row-wise (column-wise) can be decomposed and applied across individual horizontal (vertical) partitions, followed by a stitching phase that re-composes the dataframe partitions back together vertically (horizontally) in order.

Operator decomposition is simple at its core, but dataframes have unique properties that make parallelism more challenging, like logical order and mixed types in a column. Column types may change in the decomposed dataframes in unpredictable ways, requiring expensive coordination. Perhaps the most challenging aspect of dataframes is that row-wise parallelism may be followed by column-wise parallelism, so traditionally simple optimizations are not always possible. If you’re curious to learn more about the formal dataframe data model and algebra, we published our findings in VLDB2021. To learn more about the math underlying the decomposition rules, see our research paper published in VLDB 2022.

Implementation and Impact
With the what and the how figured out, we built Modin: a dataframe implementation with a theoretical foundation that can accelerate the entire pandas API. Using decomposition rules, Modin can efficiently leverage multiple cores on a machine or on a cluster to parallelize all pandas operations – leading to multiple orders of magnitude speedup. In addition, Modin’s implementation is efficient, avoiding unnecessary memory copies and leveraging compression. Modin also supports out-of-core processing, allowing performance to gracefully degrade when out of memory.

This impact on performance means that data processing notebooks and scripts in pandas that were once written as “one-offs” to be tested on small samples can now directly operate on data at scale – with no changes (see figure above). This leads to substantial productivity gains for data scientists, as well as for data engineers who are no longer required to help retranslate pandas scripts into a different framework, such as SQL, in order to run at scale.
Our experience building Modin has been incredibly rewarding– we see Modin adoption across a wide variety of sectors, including healthcare, telecommunications, finance, AI, hardware and automotive industries. From building financial Monte-Carlo simulations using pandas window functions, to AI feature extraction from documents using pandas `apply` functions, to health-care patient cross-tab diagnostics via pandas `pivots`, we’ve been delighted to see the wide variety of use-cases that Modin is empowering at scale.
Are you eager to try Modin out to improve your pandas scalability – and your team’s productivity? We’d love to help! Book a meeting with us here!
If you’d like to help us continue to develop Modin and work on some of the most interesting, impactful, and challenging data systems problems, check out our job postings here!




